Ampure Xp Beads Size Selection Chart
Ampure Xp Beads Size Selection Chart - Web size select the small rna library using ampure xp beads after using column purification. Converting ng/µl to nm when calculating dsdna library concentration. Agencourt ampure xp beads) provides the best selection for dna fragments >2 kb, and improves the median read. This cut‐off can be increased or decreased (e.g., up to 300 bp, down to 50 bp) by isolating the beads. Additionally, the dna concentration of the. Web considerations for choosing an illumina dna pcr free library preparation workflow. Small molecular species such as free adapters and adapter. We suggest using the spriselect reagent , as it is developed. Web agencourt ampure xp utilizes an optimized buffer to selectively bind dna fragments 100 bp and larger to paramagnetic beads. Warm the ampure beads to room temperature and mix thoroughly before use. Web size selection using ampure xp beads does not remove small fragments. Excess primers, nucleotides, salts, and enzymes can. Web mix the total reaction volume by pipetting 10 times and incubate at rt for 1 minute or vortex for 1 minute at an appropriate speed until homogenous (depending on labware. Web size select the small rna library using ampure xp. Web size select the small rna library using ampure xp beads after using column purification. It allows you to perform. We suggest using the spriselect reagent , as it is developed. Agencourt ampure xp beads) provides the best selection for dna fragments >2 kb, and improves the median read. Additionally, the dna concentration of the. Web can i perform size selection with the ampure xp reagent? It will be suitable to remove peaks >. To the purified pcr reaction (25 μl), add 32.5 μl (1.3x) of resuspended ampure xp. Web we found that a 0.7x volume of custom spri beads (e.g. Bead size selection is only recommended for samples showing no primer dimer and no. The following tables illustrate the number of pcr reactions th e agencourt ampure xp will. Web mix the total reaction volume by pipetting 10 times and incubate at rt for 1 minute or vortex for 1 minute at an appropriate speed until homogenous (depending on labware. How do i perform size selection? If you perform the qc check and your. Although size selection protocols developed by other organizations do exist, we cannot guarantee performance or support this application. Excess primers, nucleotides, salts, and enzymes can. If you perform the qc check and your sample contains adaptor dimer (127 bp peak) or. Web we found that a 0.7x volume of custom spri beads (e.g. We suggest using the spriselect reagent ,. Bead size selection is only recommended for samples showing no primer dimer and no adaptor dimer on bioanalyzer. Web agencourt ampure xp utilizes an optimized buffer to selectively bind dna fragments 100 bp and larger to paramagnetic beads. It will be suitable to remove peaks >. Web size select the small rna library using ampure xp beads after using column. It will be suitable to remove peaks >. Additionally, the dna concentration of the. It allows you to perform. Web size selection using ampure xp beads does not remove small fragments. Although size selection protocols developed by other organizations do exist, we cannot guarantee performance or support this application. Web size selection using ampure xp beads does not remove small fragments. If you perform the qc check and your sample contains adaptor dimer (127 bp peak) or. We suggest using the spriselect reagent , as it is developed. This cut‐off can be increased or decreased (e.g., up to 300 bp, down to 50 bp) by isolating the beads. Web. Web we found that a 0.7x volume of custom spri beads (e.g. Bead size selection is only recommended for samples showing no primer dimer and no adaptor dimer on bioanalyzer. This cut‐off can be increased or decreased (e.g., up to 300 bp, down to 50 bp) by isolating the beads. Web considerations for choosing an illumina dna pcr free library. Web considerations for choosing an illumina dna pcr free library preparation workflow. It is recommended to choose the appropriate method based on the qc check of the library. This cut‐off can be increased or decreased (e.g., up to 300 bp, down to 50 bp) by isolating the beads. Web size selection using ampure xp beads does not remove small fragments.. It is recommended to choose the appropriate method based on the qc check of the library. We suggest using the spriselect reagent , as it is developed. Additionally, the dna concentration of the. Web size selection using ampure xp beads does not remove small fragments. Web standard ampure xp is used to remove dna fragments smaller than 100 bp. Warm the ampure beads to room temperature and mix thoroughly before use. Web mix the total reaction volume by pipetting 10 times and incubate at rt for 1 minute or vortex for 1 minute at an appropriate speed until homogenous (depending on labware. Excess primers, nucleotides, salts, and enzymes can. To the purified pcr reaction (25 μl), add 32.5 μl (1.3x) of resuspended ampure xp. Web we found that a 0.7x volume of custom spri beads (e.g. Web size select the small rna library using ampure xp beads after using column purification. Agencourt ampure xp beads) provides the best selection for dna fragments >2 kb, and improves the median read. Converting ng/µl to nm when calculating dsdna library concentration. How do i perform size selection? Web the agencourt ampure xp can be used for pcr puri fication in 96 and 384 well format. If you perform the qc check and your sample contains adaptor dimer (127 bp peak) or.
Ampure XP Labplan
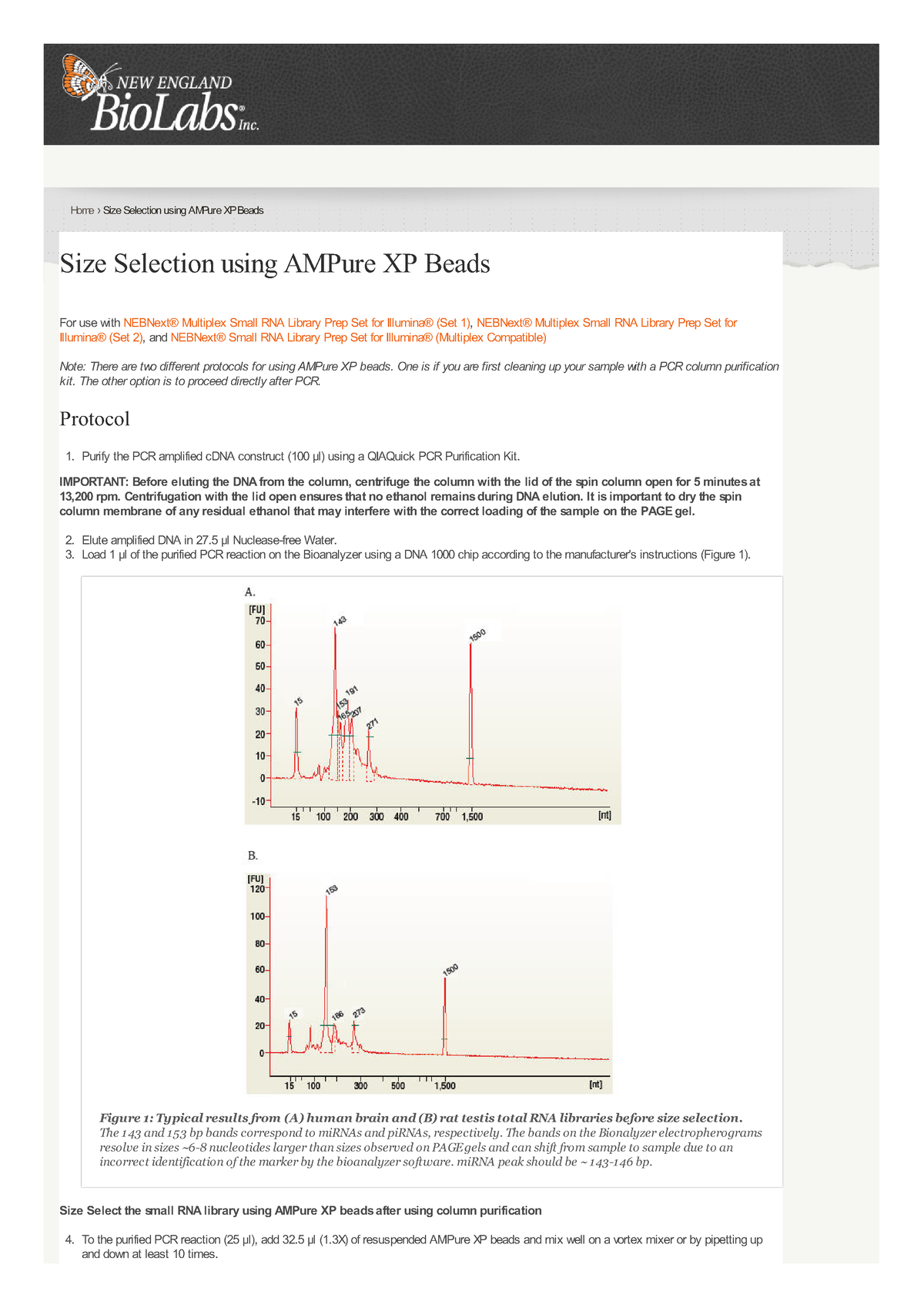
Size Selection using AMPure XP Beads Accounting Studocu

Optimising library size selection for iCLIP2 SampletoProNex bead
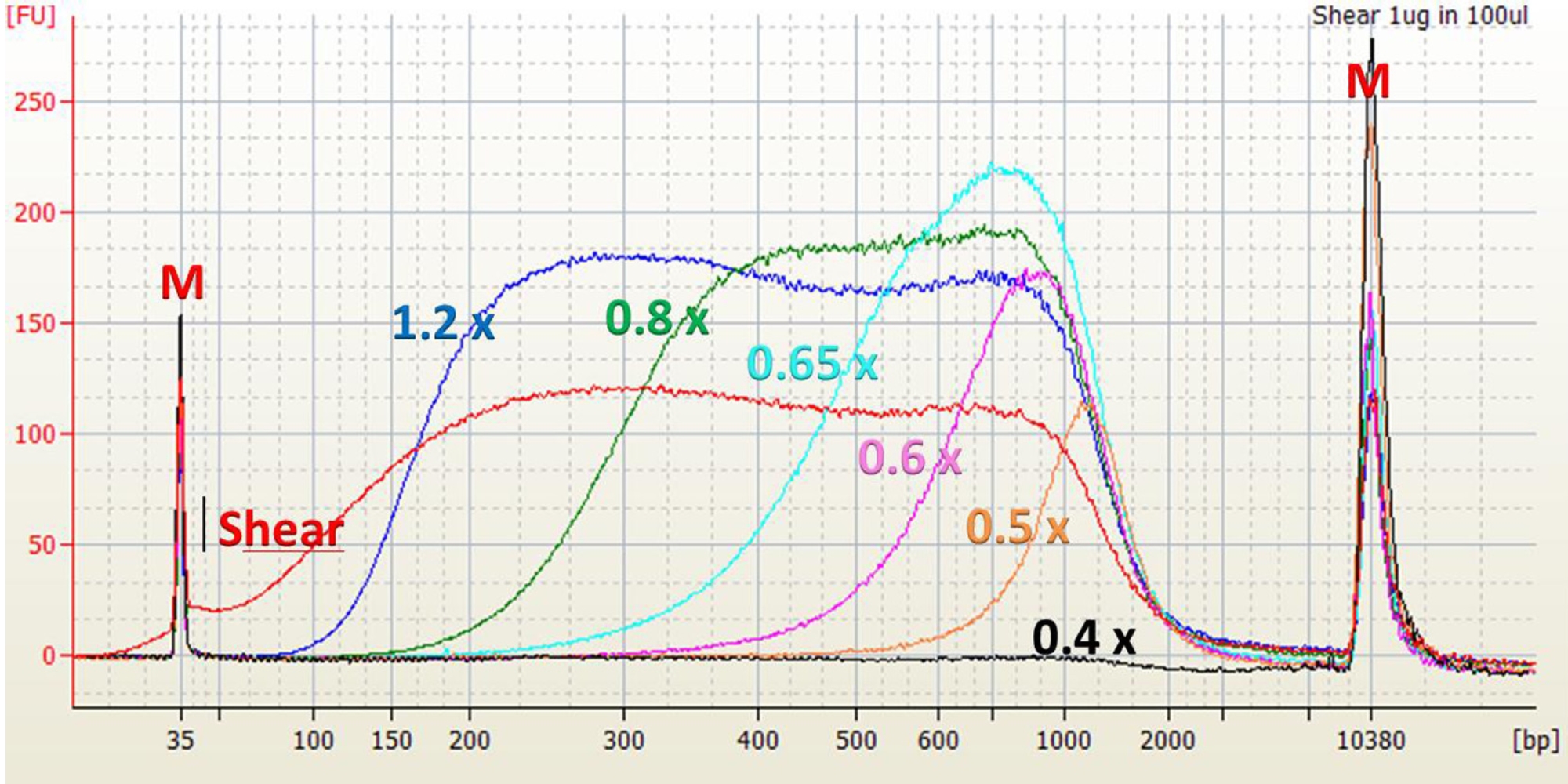
Ampure Xp Beads Size Selection Chart

Ampure Beads Size Selection Chart

AMPure XP Beads Cleanup and Size Selection

How do SPRI beads work? Enseqlopedia

Möglichkeiten mit AMPure XP Beckman Coulter

Ampure XP Labplan

Bead Size Selection and Ratio Beckman Coulter
Web Agencourt Ampure Xp Utilizes An Optimized Buffer To Selectively Bind Dna Fragments 100 Bp And Larger To Paramagnetic Beads.
Small Molecular Species Such As Free Adapters And Adapter.
Web Can I Perform Size Selection With The Ampure Xp Reagent?
Bead Size Selection Is Only Recommended For Samples Showing No Primer Dimer And No Adaptor Dimer On Bioanalyzer.
Related Post: